UF engineers reach semi-finals in XPRIZE Contest for new COVID-19 test methods; their CRISPR-ENHANCE methodology published in Nature Communications journal
The Challenge
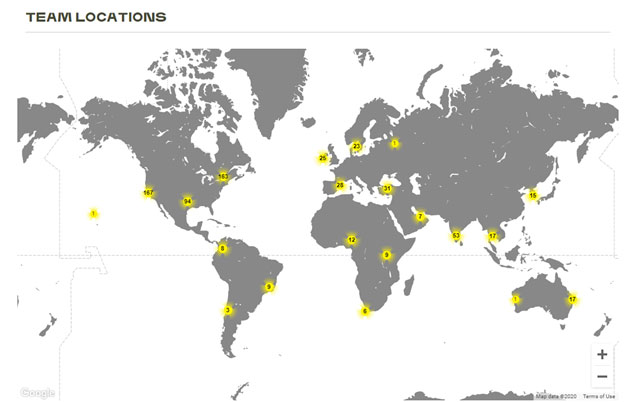
XPRIZE Rapid COVID Testing competition teams are located around the globe.
According to a survey conducted in July 2020 by researchers from Northeastern, Harvard, Rutgers, and Northwestern universities, testing (and especially testing asymptomatic individuals) is a cornerstone of containing the spread of COVID-19. The speed in producing reliable results is of the essence in controlling the spread of the virus.
Piyush Jain, Ph.D., Assistant Professor in the Department of Chemical Engineering at UF Herbert Wertheim College of Engineering, has been improving existing methods for testing COVID-19 samples from nasal swabs and saliva since the deployment of the original qPCR-based test in February 2020. His research has gained recognition by XPRIZE Foundation, Inc., a non-profit corporation fostering and sponsoring competitions to create innovative breakthroughs for the benefit of humanity.
Dr. Jain and his colleagues were selected from more than 700 teams around the globe as one of the 219 semi-finalists from 35 countries in the XPRIZE Rapid COVID Testing competition. The goal of XPRIZE Rapid COVID Testing is “to develop innovative, scalable COVID-19 testing solutions that will radically change the world by providing much needed insights to help society safely reopen – and not in some distant future, but right now.”
The New Methodology
Jain and his doctoral student Long Nguyen have developed an enhanced CRISPR/Cas (Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-associated proteins) system that amplifies the detection of the SARS-CoV-2 virus during the test process. This unique system, CRISPR-ENHANCE, shortens the time it takes to run the test by half and provides increased sensitivity for the results.
The gold-standard CDC-recommended qPCR-based tests currently in use are a fluorescence-based assay that requires an expensive thermocycler that cycles between temperatures and measures fluorescence using a camera. In contrast, Jain and Nguyen have been working on a lab-based fluorescence assay as well as a paper strip-based assay that can potentially be used for home-based tests that can significantly reduce cost and time.
“A version of our lab-based test in development uses only common lab equipment including a heater, vortex mixer, micro-centrifuge, pipettes, and a fluorescence plate reader,” Jain said.
This high-throughput lab-based test using an automated liquid handler robot is designed to analyze about 190 samples at once in just over two hours. The Jain lab is automating the entire workflow to reduce time for sample processing. With automation, pooling of samples and multiple cycles of tests in parallel, a small diagnostic lab can analyze thousands of samples daily.
If used for a single sample analysis, Jain’s entire lab-based assay can be performed in about 45-60 minutes, with the results being read in the last 5-10 minutes. In contrast, the current CDC-recommended qPCR-based tests take a total of 110-120 minutes, with the result-reading portion requiring approximately 80 minutes on the expensive thermocycler. This occupancy time becomes crucial when it comes to testing thousands of samples.
The CRISPR-ENHANCE method is also much more rapid and sensitive than two other CRISPR methods now on the market as it uses an engineered version of CRISPR that enhances the speed of CRISPR, shortening the time to receiving results by 3.2-fold. Also, if the CRISPR-ENHANCE assay runs longer, it enhances the sensitivity by several fold. The increase in cost of reagents using CRISPR-ENHANCE vs. conventional CRISPR is within $0.01 per sample, according to Jain.
Regarding the SARS-CoV-2 RNA extraction procedure, which has to occur before the reading phase, Jain said, “The CDC recommended RNA extraction/purification steps take 30-40 minutes of laborious work and use an expensive kit with supply chain issues. However, our CRISPR-ENHANCE test can potentially work with a heat-based method that employs a simple 10- to 17-minute heating step without the need for further manipulations. The cost of the extraction reagents is 80-times lower than the current CDC-approved method, with the added benefit of helping to minimize supply chain issues. We are using this approach in the XPRIZE competition.”
Next steps
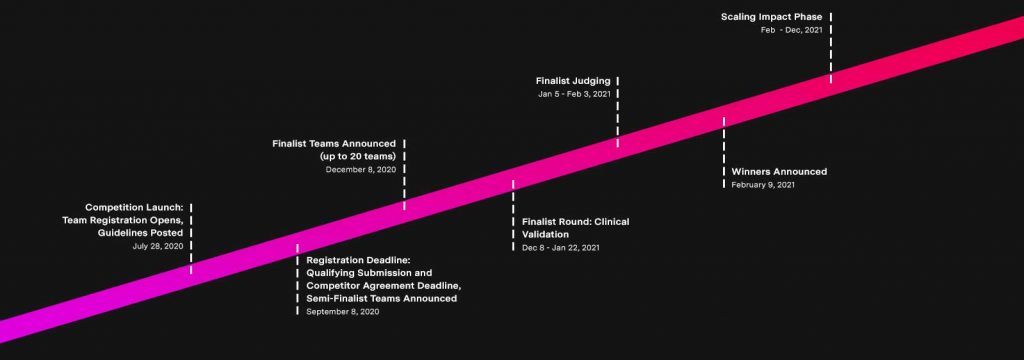
Jain’s CRISPR-ENHANCE team has tested 157 blind samples supplied by the XPRIZE Foundation and has sent those results to the judges, who will score them for specificity, sensitivity and limits of detection. From these scores, the XPRIZE Foundation will announce 20 teams on December 8, 2020, as finalists who will then have the opportunity to gain clinical validation from two world-class laboratories based in the United States, helping to accelerate the FDA approval process for their tests.
The five top testing protocols selected by the judges will be documented in a “playbook” for global dissemination in order to aid testing sites around the world in a broad effort to end the pandemic.
Nature Communications, doi:10.1038/s41467-020-18615-1, has recently published a paper by Jain, Nguyen and recent graduate Brianna Smith (B.S., MAE ‘19) on the results of their research. The paper features the development of the CRISPR-ENHANCE technology and its use in COVID-19 detection, using both the lab-based fluorescence assay and the paper strip-based assay. This paper focuses on the use of synthetic genomic RNA in the testing samples.
Jain and Nguyen hope to have their validation data on human samples completed soon and are preparing a filing with the Food and Drug Administration (FDA) to have their method approved for Emergency Use Authorization (EUA).
Following FDA approval of the enhanced fluorescence-based lab assay, Jain and Nguyen hope that, by fine-tuning their paper-strip-based assay and combining it with cell phone camera, it could potentially result in a point-of-contact and home-based tests.